Projects
Detection of the geographical origin of strawberries

The problem of adulteration of strawberries has been known for some time, as it is increasingly observed that German strawberries are stretched with foreign goods. Due to the large price differences, a re-declaration offers a high financial incentive, but besides the consumers, also the honest German strawberry farmers are massively harmed.
Currently, there are no analytical methods available to the strawberry industry that allow reliable proof of origin. The commonly used stable isotope analysis does not always provide unambiguous results and can only be carried out by very specialized research facilities, which is why a broad application is not possible.
In order to protect both honest strawberry producers and consumers from fraud, alternative analytical methods for origin determination based on objective data collection are being developed. Two complementary approaches (LC-MS/MS and ICP-MS) will be used, as it is not yet clear which technology is better suited for provenance detection. In any case, chemometric fusion of the different data sets will provide a higher resolution chemical fingerprint of the respective strawberries of different origins, which may be invaluable, for example in areas that are located on the border with neighboring countries.
After successful completion of the project, the developed methods can be directly implemented in quality assurance laboratories of SMEs, in commercial laboratories and in the laboratories of governmental investigation agencies. The project is designed so that implementation can be done with typical laboratory equipment or requires little additional investment. Alternatively, companies have the option of outsourcing the analyses to contract laboratories.
This project is funded by the German Federal Ministry of Economic Affairs and Climate Action (via AiF) through the Research Association of the German Food Industry (FEI) under project number AiF 22909 N and is being worked on by Kim Brettschneider in collaboration with Markus Fischer's research group.
Authentication of food with mobile NIR spectroscopy

In the global food supply chain, fast and reliable methods for quality and incoming goods inspection are becoming increasingly important. In particular, there is a need for inspection methods that not only provide accurate results in a short period of time, but are also inexpensive, easy to use and can be operated with minimal technical equipment. In recent years, a number of handheld instruments have been developed that meet these criteria by using near-infrared (NIR) spectroscopy. The resulting NIR spectra are characterized by superimposed signals from a large number of different molecules and are evaluated using multivariate methods as a fingerprint of the composition of the sample under investigation.
The aim of this project is to apply this approach to the analysis of different foods in order to develop rapid and simple authentication methods. A wide range of different foods will be studied. The analysis of nuts, i.e. high-fat and high-protein plant foods, which has already begun, will be extended to the analysis of high-fat and high-protein animal foods like fish. One of the aims is to analyze whether differences in the fermentation of matjes can be detected. This is important because fermentation is time consuming and costly, but also crucial to the quality of the matjes. As is often the case when applying NIR spectroscopy to aqueous foods, one challenge could be the superimposition of relevant spectral information by intense water bands. One class of food where this problem does not arise is spices, as they are usually sold in dried form. The topics investigated include the detection of turmeric in ground saffron and the addition of cassia cinnamon to Ceylon cinnamon.
As NIR spectra are used as a fingerprint of the biological samples, another aim of the research project is to deepen the understanding of the spectra generated in the context of food analysis. By fusing the NIR data with mass spectrometry and nuclear magnetic resonance spectroscopy data, a deeper understanding will be gained of the information captured by the mobile NIR instruments and how this contributes to the classification of food.
Nicolas Lauer is working on this project.
Surface-enhanced Raman scattering
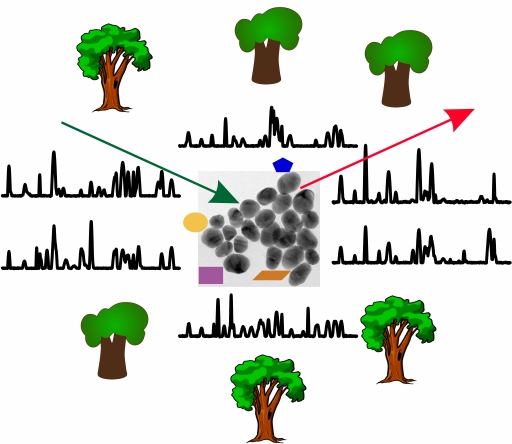
Surface-enhanced Raman scattering (SERS) in the absence of labels, tags or reporters is a powerful method to obtain comprehensive and diverse information about the composition and structure of biomolecular samples. Because of the nature of SERS that probes the varying interaction of the molecules with a nanostructured metal substrate and the proximity of specific functional groups, it is difficult to exploit the highly complex data that are generated. Specific challenges are posed by biological samples that are characterized by many different biomolecules and changing conditions, influencing the obtained SERS spectra. This is why efficient and unbiased approaches for the utilization of SERS data are needed.
In this project, random forest (RF) based methods, which have been shown to be very robust when applied to real SERS data, will be adapted and used for the analysis of model data obtained under well-defined experimental conditions. The complexity of the systems that are analyzed here will be increased step-wise from individual molecular components such as building blocks of lipid membranes or a drug molecule, over the combination of two components, to the complex environment of an endolysosomal vesicle inside cultured cells. The RF analysis will be established in such a way that it can serve three different purposes: (i) the selection of important spectral features for direct structural interpretation, (ii) the identification of co-occurring spectral features to analyze the interaction of different molecules, and (iii) the integration of a priori knowledge from previous experiments, including those conducted with respective other models in this project. The development of such a multi-purpose framework will rely on the different degrees of complexity of the SERS experiments as well as on a set of experimental conditions that are defined and modified in a systematic way. As a further aspect, simulated data that are specifically generated to imitate the impact of a particular experimental condition will be compared with actual experiments in an iterative fashion. The successful completion of this project will mark an important step towards utilizing SERS for the structural characterization of well-defined biophysical models as well as of molecules in complex biological systems.
This project is funded by the DFG under project number 511107129 and is worked on by Florian Gärber in cooperation with the research group of Janina Kneipp at the Humboldt University Berlin.
Fusion of spectroscopic and spectrometric data
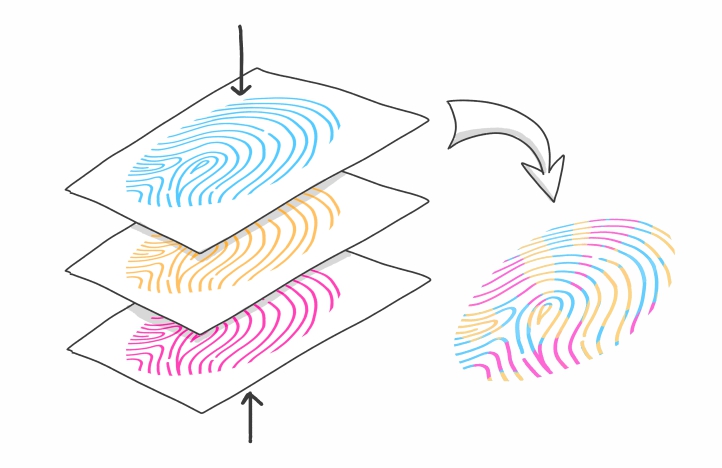
Various methods exist for generating analytical fingerprints that can be used to classify and characterise biological samples according to their origin or biological identity. These methods are often based on nuclear magnetic resonance, Fourier transform near-infrared spectroscopy, or mass spectrometry (e.g. liquid chromatography coupled with mass spectrometry, or inductively coupled plasma mass spectrometry). However, the data obtained using these methods only represent part of the samples' complex composition. This means that the chemometric models created to classify the samples using the respective data sets are also based on a limited number of existing differences.
To obtain more robust and powerful classification models, this DFG-funded project will develop and compare data fusion approaches that combine complementary information from various techniques. We will simulate model data sets with specific properties and use existing data sets of various foods. This project will therefore provide crucial methodological insights into combining molecular and elemental fingerprints, and represent an important step towards improving food authentication and detecting food fraud.
This project is funded by the DFG under project number 561375600 and is worked on by Edward Ho.
