Research Group Seifert

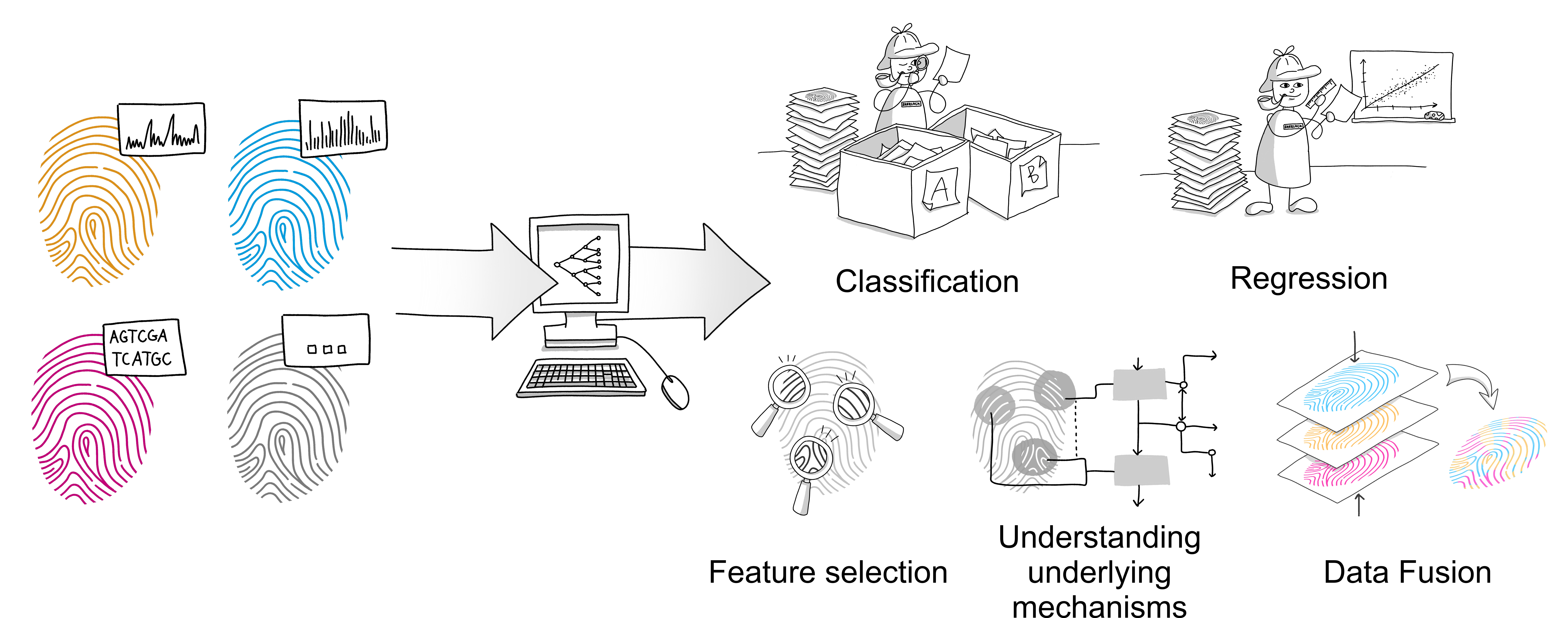
The working group analyses data from various analytical methods, which can be regarded as fingerprints of the samples examined. We are working at the interface of method development/validation of chemometric and bioinformatic approaches, mainly performed with simulated data, and their practical applications in various research areas. Of particular interest are omics data sets, which are high-dimensional data derived from the analysis of an entire cellular analyte group, thereby representing its entirety. We are also very interested in the analysis of various spectroscopic data, specifically surface-enhanced Raman scattering, e.g. to study the influence of drugs in living cells. Our analyses aim at the characterization of the samples beyond their black box classification. This means that important variables are selected and interactions of different variables are analyzed to assess the properties of the sample in as much detail as possible.
The objectives of the data analysis and the various areas of the group's work are presented below. (Graphics: freisinn.net)