Research Focus
Molecular Modelling and Medicinal Chemistry (Life Science)Understanding the molecular mecanisms of protein-ligand interactions is essential for the development of new drugs. This is the basis for identifying structure-activity relationships and resistance mechanisms on molecular level that need to be considered when designing new active ingredients. In this area, different projects are undertaken. For exploring the interaction of the inhibitors of plasmodic dihydrololate reductase (DHFR) with the target enzyme we have used information based on data obtained from the x-ray crystallography of the DHFR structures in different species to develop a homology model of the enzyme (fig. 1).
This model enables us to study the protein-ligand interaction in detail, showing possibilities to avoid the development of drug resistance in the drug development process (Lemcke, T., Christensen, I. T., Joergensen, F. S. (1999). Towards an understanding of drug resistance in Malaria: Three-dimensional structure of Plasmodium falciparum dihydrofolate reductase by homology building. Bioorganic & Medicinal Chemistry 7, 1003-1011). The model helped to explain and understand the drug resistance caused by point mutations in the plasmodic enzyme. (Modeling study carried out in the research group of Flemming Steen Joergensen, University of Copenhagen/Denmark). The enzyme class of kinases plays a significant role in the signal transmission in the cell and cell division. A large number of metabolic processes is affected or controlled by kinases. In some diseases, kinases are overexpressed (for instance in some tumors) or hyperactive (Alzheimer's disease). Therefore selective kinase inhibitors are considered very attractive targets in the development of new drugs against various diseases (cancer, diabetes, Alzheimer's disease etc.) In pursuit of selective inhibitors of different kinases (such as CDK1, CDK5, GSK-3b, c-MET), methods derived from molecular modeling such as docking (fig. 2) 3D-QSAR and virtual screening are used. Thus for paullone 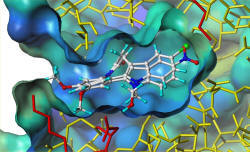 a group of new kinase inhibitors, quantitative structure-activity relationships have been calculated through comparative molecular similarity index analysis (CoMSIA) for CDK1, CDK5 and GSK-3b (Kunick, C., Lauenroth, K., Wieking, K., Xie, X., Schultz, C., Gussio, R., Zaharevitz, D., Leost, M., Meijer, L., Weber, A., Joergensen, F. S. and Lemcke, T.*; Evaluation and Comparison of 3D-QSAR CoMSIA Models for CDK1, CDK5, and GSK-3 Inhibition by Paullones. J. Med. Chem. 2004, 47, 22-36. It will be possible to interpret derived CoMSIA fields with respect to the molecular properties of new inhibitors, thus enabling the prediction of the activity of prospective-new inhibitors. In cooperation with the group of Matthias Rarey (ZBH) and Conrad Kunick (Pharmacy, University of Braunschweig) we performed a virtual screening in order to find selective inhibitors of c-Met kinase. The hits that had been found were examined by molecular modelling and synthetic experiments, thus developing new lead structures for inhibiting the c-Met kinase. In search of new drug substances for fighting malaria, the plasmodic GSK-3 has been identified as a potential molecular target for new drugs against malaria. First a homology model based on the structure of the homologous human GSK-3b was developed (Kruggel, S.; Lemcke, T.; Generation and Evaluation of a Homology Model of PfGSK-3. Arch. Pharm. Chem. Life Sci. 2009, 342, 327 – 332). Aided by this model, a structure based design is carried out to improve the activity and selectivity of existing inhibitors. With the help of molecular interaction fields (MIFs) and docking experiments, proposals for the binding mode in the ATP binding-site of the enzyme were obtained and further developed. |

 fig. 1
fig. 1