Research
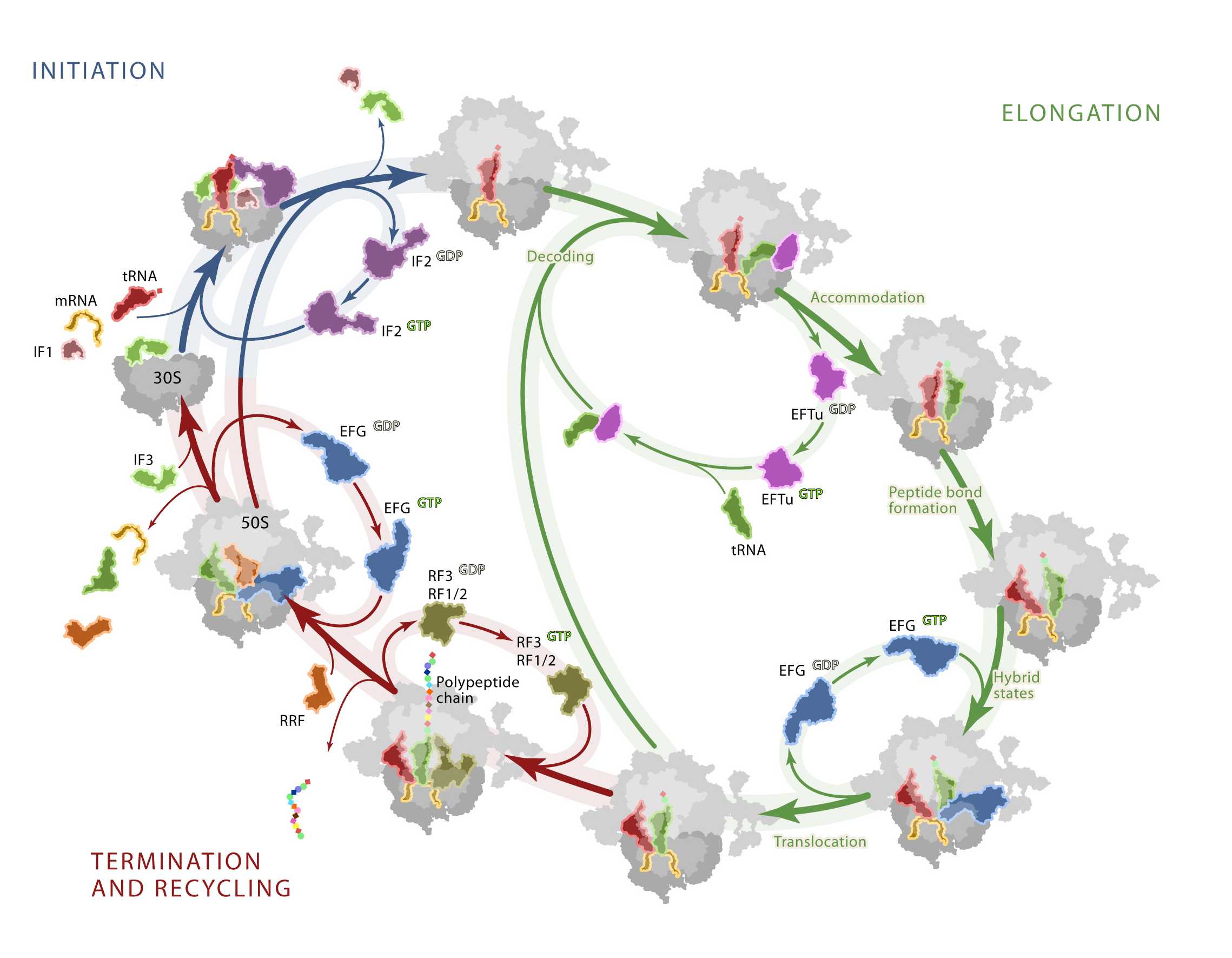
“In every living organism, translation of the genetic code into functional proteins is performed on the ribosome. Indeed, the fundamental importance of translation is reflected by the fact that the ribosome is a major target in the cell for natural antibiotics. The Wilson lab takes a combined structure-function approach to address how specific ligands, such as antibiotics as well as protein factors, modulate gene expression via interaction with the translational apparatus.”
KEY PUBLICATIONS
Structural basis of ribosomal 30S subunit degradation by RNase R
Lyudmila Dimitrova-Paternoga, Sergo Kasvandik, Bertrand Beckert, Sander Granneman, Tanel Tenson, Daniel N. Wilson , Helge Paternoga
Nature 626, 1133-1140 (2024)
https://www.nature.com/articles/s41586-024-07027-6
Structural conservation of antibiotic interaction with ribosomes
Helge Paternoga , Caillan Crowe-McAuliffe, Lars V. Bock, Timm O. Koller , Martino Morici, Bertrand Beckert, Alexander G. Myasnikov, Helmut Grubmüller, Jirí Novácek, Daniel N. Wilson
Nature Structural & Molecular Biology 30, 1380-1392 (2023)
https://www.nature.com/articles/s41594-023-01047-y
Structure of Gcn1 bound to stalled and colliding 80S ribosomes
Agnieszka A. Pochopien, Bertrand Beckert, Sergo Kasvandik, Otto Berninghausen, Roland Beckmann, Tanel Tenson, Daniel N. Wilson
Proceedings of the National Academy of Sciences of the United States of America 118, e2022756118 (8 pp) (2021)
https://www.pnas.org/doi/full/10.1073/pnas.2022756118